“Variant Normalization”的版本间的差异
小 (已保护“Variant Normalization”([编辑=仅允许管理员](无限期)[移动=仅允许管理员](无限期))) |
Suqingdong(讨论 | 贡献) |
||
| (未显示1个用户的8个中间版本) | |||
| 第37行: | 第37行: | ||
<h1>hello world</h1> | <h1>hello world</h1> | ||
| + | |||
| + | == 图片引用 == | ||
| + | [[文件:PAH.png]] | ||
== 参考 == | == 参考 == | ||
| − | + | * https://www.liwei8090.com/10586.html | |
| + | * https://genome.sph.umich.edu/wiki/Variant_Normalization | ||
| + | |||
| + | [[category:常见问题]] | ||
| + | |||
| + | ---- | ||
| + | {{SQD的模板}} | ||
| + | |||
| + | <comments /> | ||
2020年1月18日 (六) 09:39的最新版本
目录
Introduction
The Variant Call Format (VCF) is a flexible file format specification that allows us to represent many different variant types ranging from SNPs, indels to copy number variations. However, variant representation in VCF is non-unique for variants that have explicitly expressed reference and alternate sequences. A failure to recognize this will frequently result in inaccurate analyses.
On this wiki page, we describe a variant normalization procedure that is well defined for biallelic as well as multiallelic variants. We then provide a formal proof the procedure's correctness.
Definition
The normalization of a variant representation in VCF consists of two parts: parsimony and left alignment pertaining to the nature of a variant's length and position respectively.
Parsimony
In the context of variant representation, parsimony means representing a variant in as few nucleotides as possible without reducing the length of any allele to 0. It is a property describing the nature of the length of a variant's alleles and is defined as follows:
A variant is parsimonious if and only if it is represented in as few nucleotides as possible an allele of length 0.
测试边框
- solid 单线边框
- border:1px solid #808080
常用边框之一,推荐
- dashed 虚线边框
- border:1px dashed #808080
常用边框之一,推荐
- double 双线边框
- border:3px double #808080
常用双线边框之一,推荐
测试
hello world
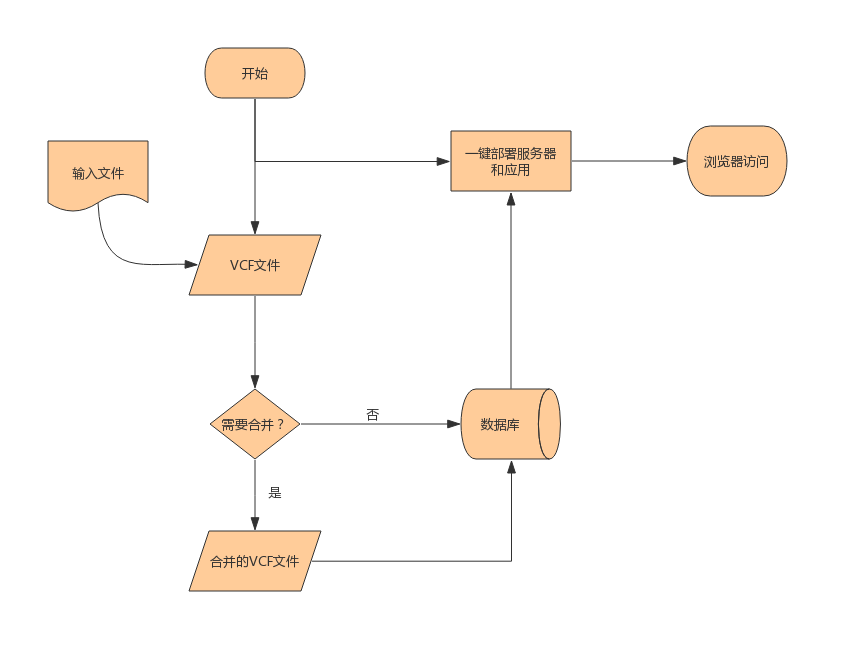
图片引用
参考
| 用户留言: |
为什么要对VCF进行norm操作?norm是针对INDEL不同表示形式的统一 |
| 新增留言 编辑留言 |
开启评论自动刷新